|
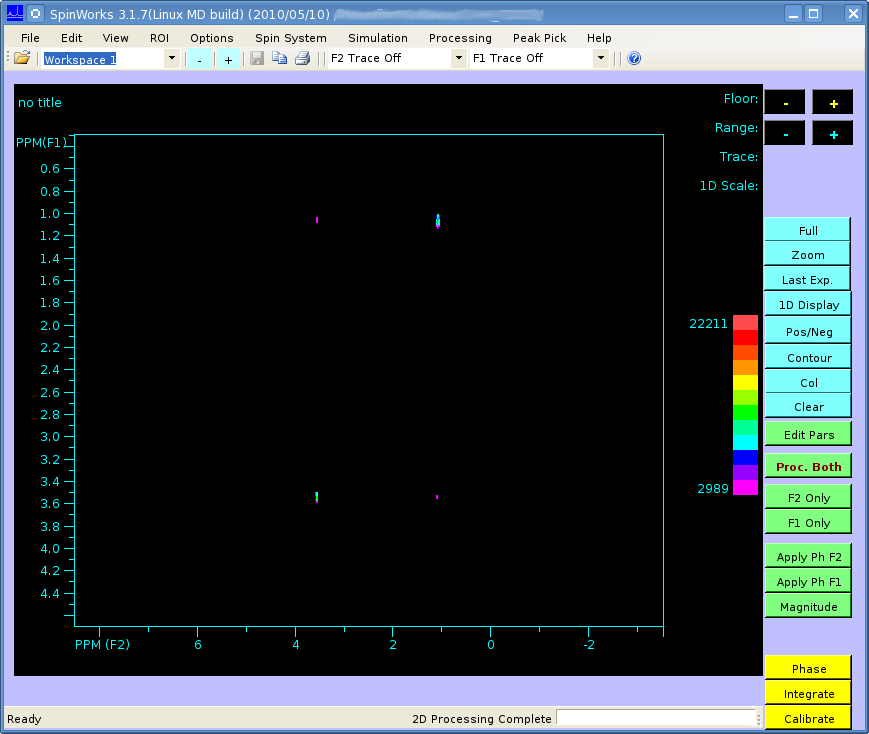
Hi, I think this is probably a basic problem but I'm a bit inexperienced with NMR. I'm just trying to process an ethanol TOCSY spectrum from the biomolecular magnetic resonance data bank, but strange peaks are appearing at the very edge of the data. The spectrum should look like this (processed in SpinWorks):
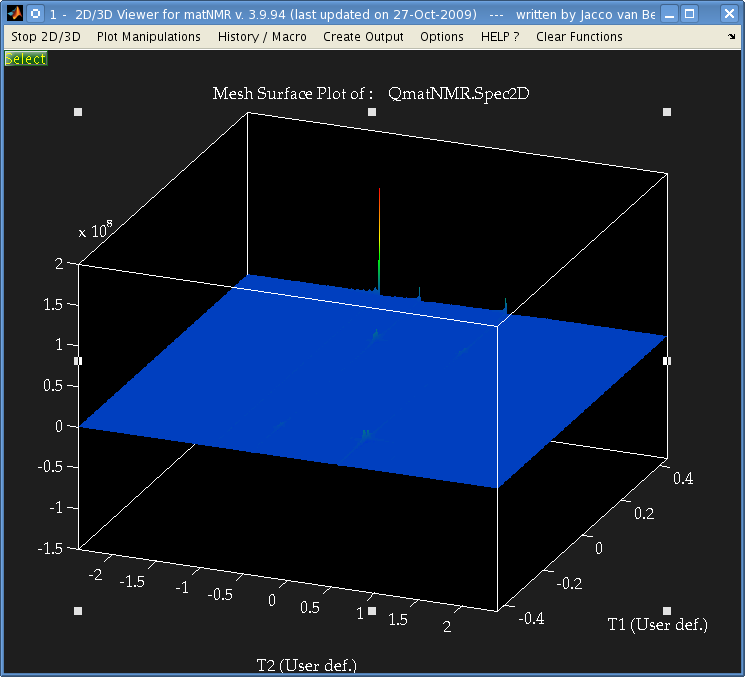
But when I perform a 2-dimensional Fourier transform using matNMR, or the fft Matlab functions I get the following:
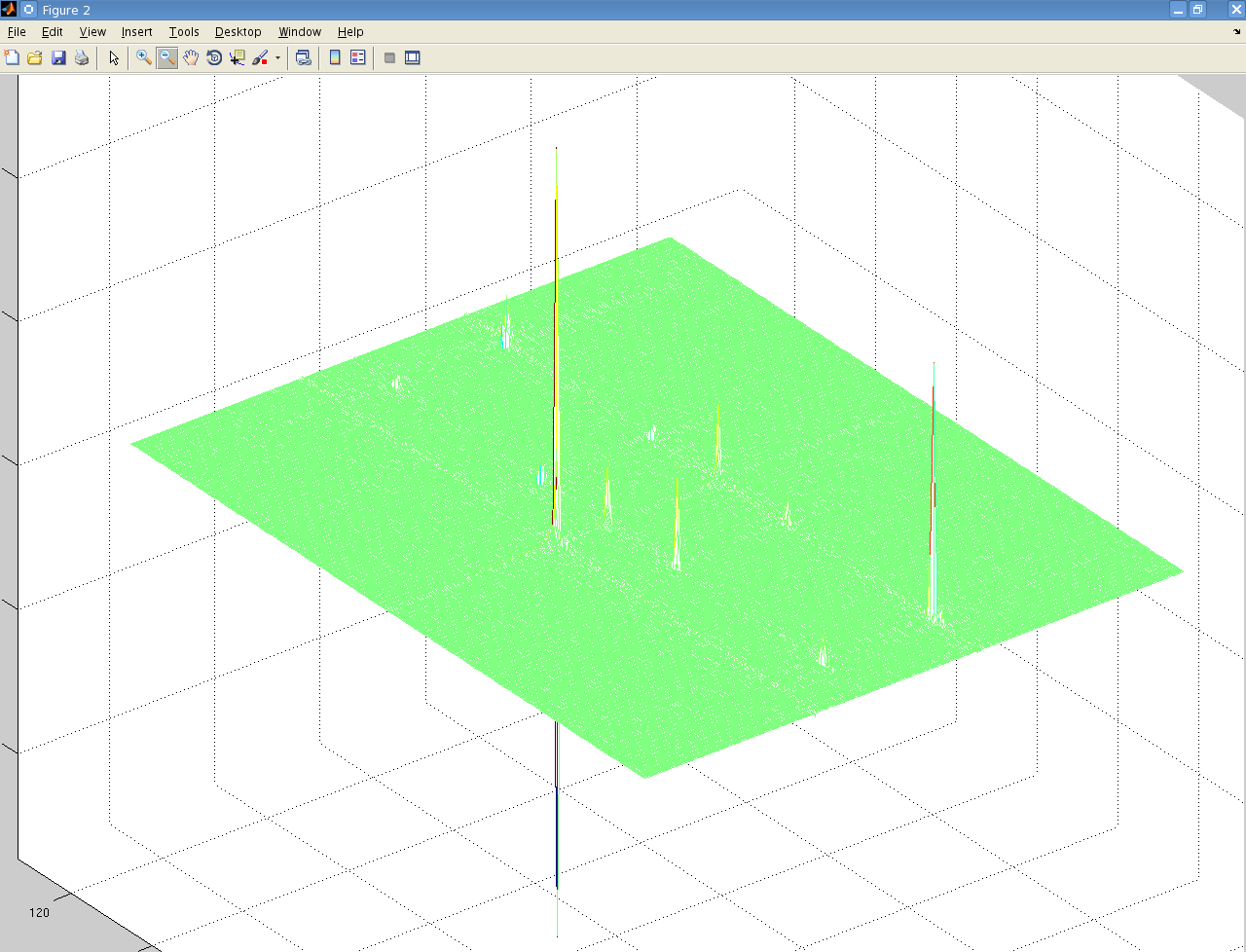
It's not easily visible but there are 4 small peaks in the body of the spectrum as well as the 3 large ones at the very edge. The data is from a Bruker spectrometer recorded using TPPI. There were zeros at the start of the FID so I used the "remove Bruker digital filter" option before processing. I performed a Fourier Transform in the T2 direction, and then T1 direction and that is the result. I don't understand why there are peaks at the very far edge of the data, and performing an fftshift results in 3 peaks at zero frequency. Is this a result of a problem with the phase, or possibly because of solvent peaks? I used the automatic phase adjustment but the peaks at the edge were still there. Is there some mistake I'm making that causes those peaks to appear at the edge? I processed data from another brand of spectrometer which had used STATES and the spectrum was as expected without anything at the edges. Also, when I process other Bruker TPPI spectra from the same data bank for adanine TOCSY and HSQC, and methanol I have the same problem. Any ideas/suggestions would be much appreciated, Thanks Update: After writing my own function to parse the ser files I got this result after taking the complex Fourier transform:
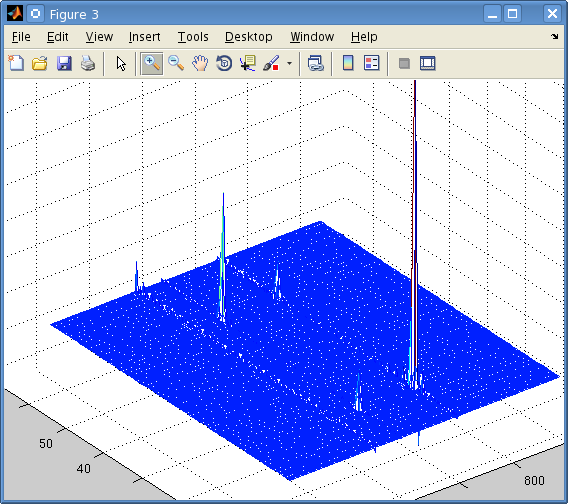
Since the complex Fourier transform of TPPI FIDs gives a symmetric spectrum, could I just truncate half of it and shift it so that 0ppm is in middle of half of the full spectrum? It seems like the 4 peaks on either side of the 3 central peaks are (kind of) symmetic? I did this and obtained the following:
I thought this seemed somewhat reasonable although was not entirely sure because the spinworks spectrum only has positive ppm values for the indirect dimension but if I shift the axes so 0ppm is the center there will be negative values, right? It was mentioned that if I wanted to take the real Fourier transform I could zero the imaginary values and take the complex Fourier transform (if I've understood correctly) but then I would need another algorithm to extract the data, is there somewhere I could read about the algorithm for extracting the data? When I did this and viewed the spectrum it seemed like there were four "copies" of the original. Also, in the process of writing the parser I came accross a few sources that mentioned the 2nd and 3rd integers in every group of 4 integers in the ser file needed to be multiplied by -1. Is there an explanation somewhere for this? I thought that simply by modifying the phase according to TPPI we could achieve quadrature detection in the indirect dimenison. Thanks again for the advice. |
|
I am not familiar with MatNMR at all. However, since your data processes O.K. in SpinWorks (and presumably in TopSpin) then the data are O.K. I suspect that the problem is that you are not handling the TPPI data correctly. You need to use a real transform in F1. One way to do this is to expand the data into the real locations only, and insert zeros into the imaginary locations. Then apply the the standard fft algorithm and slightly different algorithm to extract the data from the array after transform. I'm not sure how you would do this in MatLab. Thanks for your advice -- and sorry to reply so late. I decided to write a small parser to get data from the ser files (if nothing else, for the pedagogical value :)), and have updated my previous post with the results, there were just a few things I wasn't quite sure of which I've mentioned above. - newToNMR (Nov 12 '10 at 03:33) In the SpinWorks ftp site there is a file with three sample 2D TOCSY data sets: one recorded with States detection, one recorded with TPPI detection and another with States-TPPI detection. The file is tocsy_avance300.zip. This may help you sort things out. - Kirk Marat (Nov 15 '10 at 14:37) |